AI readiness doesn't always require a better algorithm. Sometimes, it just requires better data curation and analysis.
A case study published in PNAS makes the point. Researchers classified nearly 10,000 chemical compounds using a standard machine learning approach and achieved only 10% accuracy. When CAS re-ran the analysis with higher-quality chemical descriptors, accuracy jumped to 45% (median improvement: 33%). Same model, stronger foundation.
When the data is working harder, research teams can identify top compounds in hours instead of weeks. However, getting there has specific requirements.
[H2] When the AI you stand up falls down
There’s a story that often plays out across scientific R&D teams charged with applying AI to their research. The team:
- Invests in infrastructure.
- Hires data scientists.
- Runs pilots.
- Tests the AI systems.
- Reviews the brilliant test results.
And only then do they find out that their AI models stumble when encountering real-world conditions. Even worse? The brilliant results never arrive at all, not even within the model.
The team invests, hires, pilots, tests... and accuracy barely moves. They tweak the algorithm, add more data, adjust parameters. Still nothing. The instinct is to blame the model. But often, the real issues are upstream—and that can include fragmented data silos, inconsistent metadata, varied experimental conditions, incomplete historical records, and the lack of standardized formats across labs or instruments.
Results like that point to a gap in AI readiness.
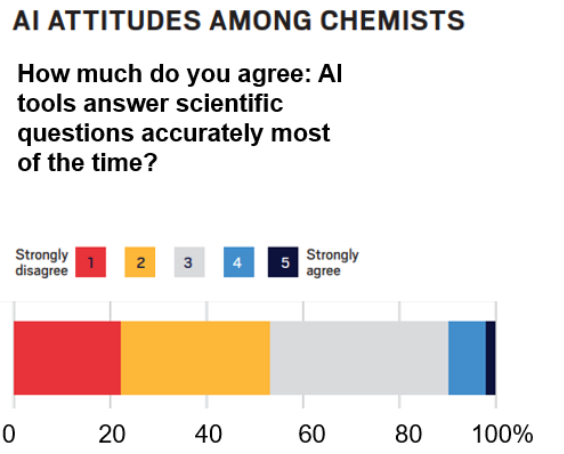
Figure 1: Source: C&EN Brand Labs Marketing for the Chemical Sciences Survey Report 2025.
While the team may strengthen their digital infrastructure, they often still struggle to make AI in R&D behave consistently in day-to-day research.
Scientific data frequently varies in structure or quality, and this variation affects how accurately algorithms can interpret the information they receive. As a result, models that perform well in controlled tests often underperform when exposed to the uneven conditions of actual experiments. For example, evaluation datasets can be “too clean” or too similar to training data. They can also fail to reflect real-world noise, outliers, experimental drift, or the full diversity of cases that researchers encounter.
Models trained on incomplete or inconsistent data never perform well in the first place. A lack of domain-specific context in AI pipelines, such as precise chemical ontologies, mechanistic constraints, or expert-curated annotations, can also cause accuracy problems that stem not from data quality itself, but from missing scientific structures. This gap introduces uncertainty, making it challenging to integrate AI into R&D workflows.
This is not just about tweaking current processes to work in a slightly different environment. Preparing to use AI as part of your core R&D infrastructure is a major undertaking. Getting it wrong can waste budgets, cause pilots to fail, discourage leadership from investing the necessary resources, and leave your research scientists skeptical about the benefits.
[H2] Three areas where organizations struggle to use AI in R&D workflows
Many organizations working to deploy AI in scientific R&D encounter similar operational roadblocks.
Most of these issues fall into a small set of recurring patterns that limit AI readiness. The following areas represent the most common sources of friction:
1. System alignment becomes a challenge when the tools that store and process scientific data cannot exchange information cleanly. This misalignment typically manifests itself in several ways:
- ELNs (Electronic Lab Notebooks) and LIMS (Laboratory Information Management Systems) often store information in incompatible formats.
- Teams must manually adjust records when data moves between systems.
- Manual adjustments slow pilot work and increase the risk of errors.
- Pipelines become unstable when metadata is recreated or renamed during transfers.
Together, these breakdowns interrupt the flow of information and prevent models from operating consistently across routine scientific tasks.
2. Manual handoffs and process interruptions erode traceability across the workflow. When data moves between systems or teams through informal channels like exports, emails, and spreadsheets, context gets lost:
- Records arrive without the metadata needed to interpret them.
- When a model produces unexpected output, teams struggle to reconstruct how the inputs were generated.
- It becomes difficult to determine whether an issue originated in the data or in a process gap.
- Investigations drag on, and scientists spend more time chasing provenance than analyzing results.
Workflow fragmentation creates communication gaps. This is especially costly when data crosses departmental lines. At Calibr-Skaggs, biology and medicinal chemistry teams were manually verifying as many as 1,000 novel compounds per day, slowing go/no-go decisions. CAS streamlined their pipeline with an API integration that let both teams work in parallel, eliminating delays caused by sequential handoffs.
3. Poor coordination across teams can weaken AI readiness over time.
Another common challenge occurs when science, data, and IT groups base their expectations for data structure, standards, and meaning on separate goals, processes and platforms. While some of these disconnects can be as simple as inconsistent rules for naming, versioning, or organizing records, other cases are a result of fundamental differences in meaning. These inconsistencies tend to surface when:
- Each group applies its own conventions for data definitions, structure, and organization.
- Differences require substantial manual effort to harmonize, slowing down data flow and introducing opportunities for error.
- AI teams lack knowledge of upstream differences in standards, resulting in inaccurate and unusable AI solutions.
Over time, these inconsistencies become more challenging to manage and impact confidence in AI outputs. We’ve observed these types of situations when experimental scientists are tasked with developing a full data strategy yet require additional education to handle various data formats. This is another way in which CAS staff work with partners such as Toray, helping to formalize data governance processes.
[H2] Five markers of AI readiness in R&D
Organizations that are well-positioned to scale AI share a consistent set of workflow characteristics. These five characteristics reflect how effective data, tools, and scientific reviews support models that can accommodate evolving experimental conditions, resolve domain-specific ambiguity, and incorporate continuous changes to underlying knowledge.
1. Data governance that directs scientific and technical decisions
Strong governance is a key marker of AI readiness, as it clarifies the process by which data and model decisions are made. In mature environments, ownership of datasets and their organizing rules are defined, and teams know who is responsible for approving changes to model behavior. These expectations are built into existing scientific workflows, ensuring that decisions occur in the appropriate context.
It’s also important to maintain a shared log for all datasets and model updates complete with timestamps and rationales to create the audit trail that structured oversight requires. Any changes that affect downstream workflows should also trigger a cross-functional review, ensuring that decisions made in one part of the pipeline don't introduce failures elsewhere.
When governance functions this way, teams can track what changed, why it changed, and how those updates affect AI performance across the R&D workflow.
2. Complete and scientifically meaningful metadata
AI-ready organizations treat metadata as the information that makes scientific records interpretable and actionable. Metadata describes how samples were prepared, how structures were represented, which identifiers were assigned, and where each record originated. High-readiness teams ensure that these details are complete and expressed consistently, so records do not conflict when they are passed between systems. They also maintain a history of how each record evolved. Stable, well-structured metadata enables AI systems to interpret inputs more accurately, reducing the uncertainty that often arises later in the workflow.
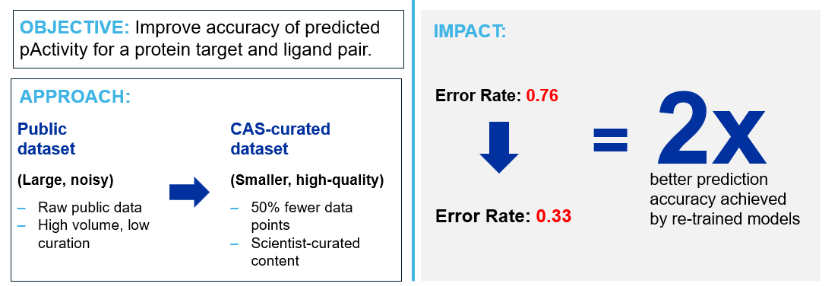

Figure 2: CAS case study showcasing the impact of high-quality, scientist-curated content on the predictive accuracy for the biological activity of a ligand against a protein target. By using CAS-curated content instead of noisy public data, the model increased 2x in prediction accuracy while using a training data set that was half the size of the original.
3. Interoperability across ELNs, LIMS, and model environments
Interoperability is another indicator of readiness, as it reflects how smoothly information flows across R&D systems. In mature environments, data is transmitted through maintained APIs rather than manual transfers, and dataset structures remain consistent as information is exchanged between ELNs, LIMS, and modeling tools. This prevents unexpected changes to fields or labels that would otherwise disrupt pipelines.
Consistent terminology across systems ensures that records retain their meaning as they move through the workflow. Solutions such as a custom API can apply metadata standards across teams to improve consistency, a strategy that has shown success with drug discovery organizations that have cross-functional teams providing data.
When interoperability functions well, models receive inputs that match real laboratory conditions, which improves stability during deployment.
4. Systems that maintain reliability as workflows evolve
Another sign of AI readiness is an organization's ability to maintain model reliability as data volume increases and scientific knowledge evolves. Mature environments maintain datasets in a versioned state, allowing earlier results to be reproduced, and they retain registries that store the data, configurations, and evaluation results associated with each model. Automated monitoring detects unexpected changes in performance, and configuration logs allow teams to understand how models were created.
These practices provide visibility into model behavior, which supports informed scientific decision-making as workflows evolve. This also ensures that scientific results and model evaluations can be reproduced months or years later, a requirement for regulatory and publication standards.
5. Human-in-the-loop validation integrated into scientific workflows
AI-ready organizations ensure that scientific review remains at the center of their workflow. Because AI cannot determine whether its predictions align with laboratory conditions, researchers review outputs at defined points to confirm that the results align with experimental expectations. Validation criteria reflect what is feasible in the laboratory and the constraints that apply to the underlying methods. These checks occur before results are passed downstream, and reviewers document the reasoning behind each decision. Regular comparison of predictions with experimental outcomes provides teams with a clearer understanding of how models behave over time and strengthens confidence in their use.
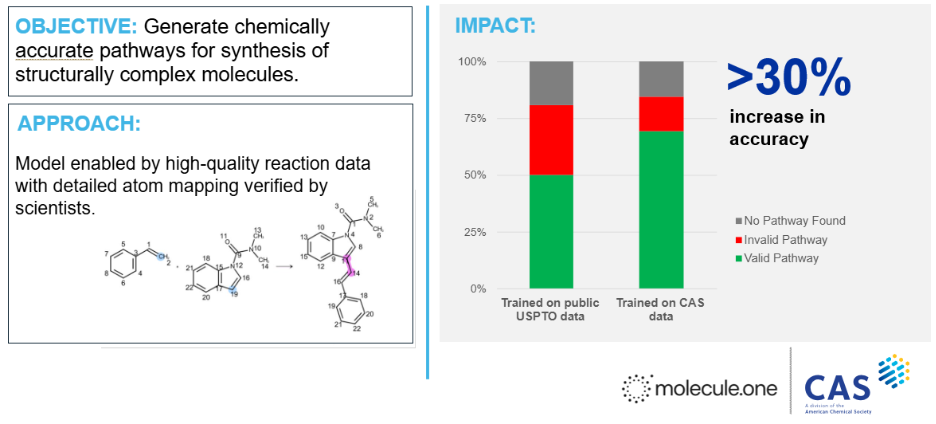
Figure 3: Case study developed in collaboration with Molecule.One. Molecule.One’s retrosynthesis model trained on public reactions data was re-trained using CAS data, leveraging the scientist-curated mappings between reactant and product atoms that are critical to accurate chemical predictions. With CAS data, accurate prediction of valid synthetic routes increased by 30%, as rated by expert synthetic chemists.
Taken together, thoughtful changes across these five areas can enhance workflow consistency, enabling AI to produce results that researchers can trust and reproduce.
[Breaker]: Research groups implementing AI strategies but uncertain about data quality and governance can consult with CAS specialists who combine scientific expertise with data infrastructure experience to assess readiness and identify gaps in existing systems.
[H2] Building the foundation for science-smart AI
When AI is ready for use in scientific R&D, its predictions give researchers information they can interpret, trust, and act on. Systems reach this point when they perform consistently under the variability of real experiments, rather than only during controlled testing. When models remain stable despite differences in instruments, formats, conditions, or workflows, AI shifts from an exploratory pilot to a practical tool that strengthens scientific decision‑making. At which point researchers can rely on AI outputs as a predictable part of their daily work, and organizations can begin scaling its use across broader R&D pipelines.
Explore how these principles translate into real R&D improvements in the CAS Science-Smart AI resources.







